Welcome to our CUT&RUN and CUT&TAG NGS Analysis Webpage!
CUT&RUN (Cleavage Under Targets & Release Using Nuclease) and CUT&TAG (Cleavage Under Targets & Tagmentation) are next-generation sequencing (NGS) techniques that enable the mapping of genome-wide protein-DNA interactions with high sensitivity and specificity. Basepair provides a point & click graphical user interface to the industry standard tools needed to analyze this type of data (such as Bowtie2, SEACR, Homer and MACS2) so you don’t have to spend time figuring out how to run them yourself. Analyze Six Samples For FreeSome of our Customers




Easy as 1, 2, 3.
Upload, analyze, visualize.
We wanted to make CUT&RUN and CUT&TAG data analysis so simple that users with no programming experience, or the time to wait for days for results, could run our automated pipelines. Data upload is easy. Just drag and drop files, connect directly to Basespace or use your own cloud storage bucket. If an interesting relationship or trend in the data is observed in one of the visualizations in our interactive reports, then maybe that’s the time to collaborate with a bioinformatician on an informed question.
Have an Active Motif kit to help generate your data? We have partnered with Active Motif to make it easy to run their approved workflows. What’s more, they’ll provide you with two free sample analysis credits when you log into their analysis portal, powered by Basepair. Just ask an Active Motif sales rep for the coupon code.


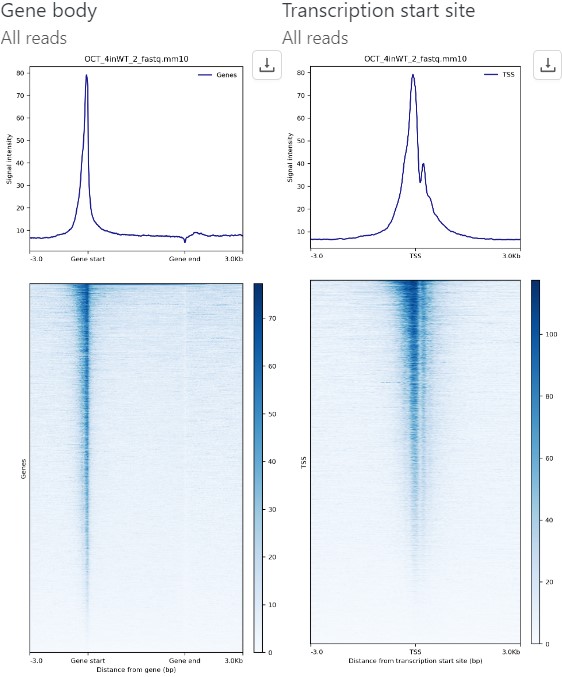
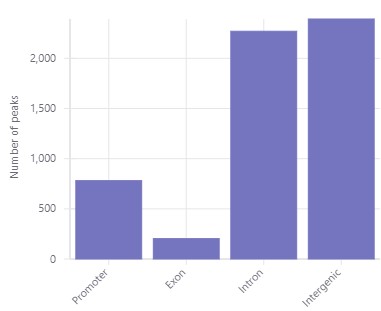
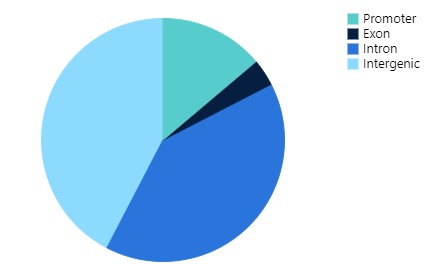
Transparency & Reproducibility with Industry Standard Tools
Our platform was designed for reproducibility and transparency — from QC and read charts, to interactive plots ready for publication. All CUT&RUN and CUT&TAG tools available in our platform are industry standard algorithms that have been highly cited and peer-reviewed. They include:
- Bowtie2 is a fast and accurate alignment tool that aligns the high-quality reads to a reference genome or assembly with high sensitivity and specificity. It is designed to handle a wide range of read lengths and is suitable for analyzing CUT&RUN and CUT&TAG NGS data from various organisms and experimental conditions.
- SEACR (Signal Extraction for Autocorrelated Regions) is a tool specifically designed for CUT&RUN and CUT&TAG NGS data analysis. It extracts the signal from the data and identifies enriched regions with high sensitivity and specificity. SEACR also corrects for the autocorrelation inherent in such data, resulting in more accurate peak calling and improved downstream analysis.
- Homer is a suite of tools for analyzing ChIP-seq as well as CUT&RUN and CUT&TAG NGS data. It includes tools for motif discovery, peak calling, and differential analysis. Homer’s peak calling algorithm uses a dynamic background model to accurately identify peaks, even in noisy data. It also includes tools for functional annotation, including gene ontology and pathway analysis.
- MACS2 (Model-based Analysis of ChIP-Seq 2) is another popular tool for peak calling in both ChIP-seq as well as CUT&RUN and CUT&TAG NGS data. It uses a probabilistic model to identify peaks and includes a number of options for peak calling and peak filtering. MACS2 also includes tools for visualization, annotation, and differential analysis.
Commercial Grade Software at an Affordable Price
Let’s face it, no one likes annual software licenses unless you need to process large sample volumes. At Basepair you can get started with as little as $15 per sample with our pay-as-you-go usage model, with no upfront license fee, making it accessible for any budget. That includes everything you need – all the tools, analysis compute, unlimited access to the visualizations and even 12 months cloud storage so you can access your data anywhere, anytime. We also offer annual plans catered to individual needs.

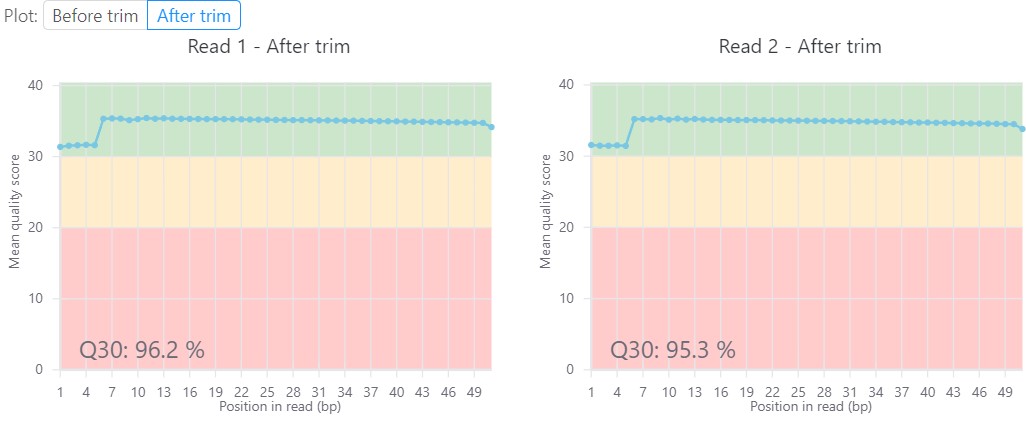
More Than Just Flat File Outputs
But if that’s what you want, all of the intermediary files are made available for download. All of our graphs and figures can also be downloaded as high quality, ready-for-publication images to make writing that paper or creating that powerpoint just a little bit easier.
Otherwise all of the interactive visualizations update dynamically as you change the parameters & thresholds, allowing you to explore your data at length to facilitate that next scientific insight.
What about my Bioinformatics team?
Despite our simplified exterior, Basepair is powered by powerful bioinformatics algorithms under the hood. We already support standard analysis pipelines and if needed will gladly collaborate with your bioinformatics team to deploy your custom, approved workflows to the platform. What’s more, Basepair is extensible through our CLI and powerful APIs so that your Bioinformatics team can interact with it in a way that they are more familiar with.

Ready to take your CUT&RUN or CUT&TAG data analysis to the next level?
Don’t take our word for it, try all of these features for yourself. Inspect finished reports, use sample data or upload your own with our free, no-commitment 14-day trial, and see why some of the world’s top institutions are using Basepair to save time and money on their NGS data analysis.
Basepair Work is in Dozens of Peer-Reviewed Journals
“Fast, excellent and reasonably priced…you CAN get all three!! Thank you to the folks at Basepair for helping us deal with some difficult RNA Seq data.”
“I really like how easy the website is to use. And how quickly the results are generated, including figures. I would have never thought about doing a new analysis like I just did.”
“Support answers come fast and are always precise!”






